Assembly statistics and evaluation¶
Note
If you are starting at this point, you’ll need a copy of the assembly we just performed (Running the actual assembly). You can set that up by doing:
sudo chmod a+rwxt /mnt
cd /mnt
mkdir work
cd work
curl -L -O https://github.com/ngs-docs/2016-aug-nonmodel-rnaseq/raw/master/files/assembly-and-reads.tar.gz
tar xzf assembly-and-reads.tar.gz
So, we now have an assembly of our reads in rna-assembly.fa. Let’s
take a look at this file –
head rna-assembly.fa
This is a FASTA file with complex (and not, on the face of it, very informative!) sequence headers, and a bunch of sequences. There are three things you might want to do with this assembly - check its quality, annotate it, and search it. Below we’re going to check its quality; other workshops do (will) cover annotation and search.
Applying transrate¶
transrate is a program for assessing RNAseq assemblies that will give you a bunch of assembly statistics, along with a few other outputs.
First, let’s download and install it:
cd
curl -O -L https://bintray.com/artifact/download/blahah/generic/transrate-1.0.3-linux-x86_64.tar.gz
tar xzf transrate-1.0.3-linux-x86_64.tar.gz
export PATH=~/transrate-1.0.3-linux-x86_64:$PATH
Now run transrate on the assembly to get some preliminary stats:
cd /mnt/work
transrate --assembly=rna-assembly.fa --output=stats
This should give you output like this:
[ INFO] 2016-08-20 10:25:51 : n seqs 60
[ INFO] 2016-08-20 10:25:51 : smallest 206
[ INFO] 2016-08-20 10:25:51 : largest 4441
[ INFO] 2016-08-20 10:25:51 : n bases 73627
[ INFO] 2016-08-20 10:25:51 : mean len 1227.12
[ INFO] 2016-08-20 10:25:51 : n under 200 0
[ INFO] 2016-08-20 10:25:51 : n over 1k 34
[ INFO] 2016-08-20 10:25:51 : n over 10k 0
[ INFO] 2016-08-20 10:25:51 : n with orf 38
[ INFO] 2016-08-20 10:25:51 : mean orf percent 67.01
[ INFO] 2016-08-20 10:25:51 : n90 816
[ INFO] 2016-08-20 10:25:51 : n70 1562
[ INFO] 2016-08-20 10:25:51 : n50 1573
[ INFO] 2016-08-20 10:25:51 : n30 2017
[ INFO] 2016-08-20 10:25:51 : n10 3417
[ INFO] 2016-08-20 10:25:51 : gc 0.47
[ INFO] 2016-08-20 10:25:51 : bases n 0
[ INFO] 2016-08-20 10:25:51 : proportion n 0.0
...which is pretty useful basic stats.
I’d suggest paying attention to:
- n seqs
- largest
- mean orf percent
and more or less ignoring the rest on a first pass.
Note: don’t use n50 to characterize your transcriptome, as with transcripts you are not necessarily aiming for the longest contigs, and isoforms will mess up your statistics in any case.
Evaluating read mapping¶
You can also use transrate to assess read mapping; this will use read evidence to detect many kinds of errors.
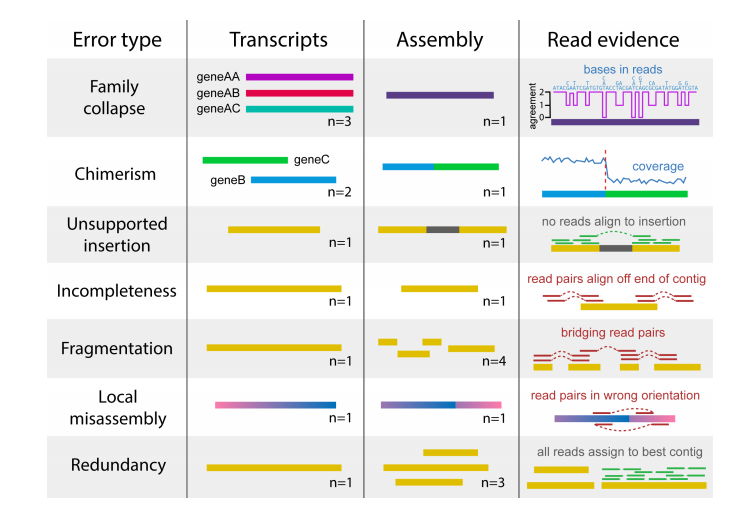
To do this, you have to supply transrate with the reads:
transrate --assembly=rna-assembly.fa --left=left.fq --right=right.fq --output=transrate_reads
The relevant output is here:
[ INFO] 2016-08-20 10:27:47 : Read mapping metrics:
[ INFO] 2016-08-20 10:27:47 : -----------------------------------
[ INFO] 2016-08-20 10:27:47 : fragments 57158
[ INFO] 2016-08-20 10:27:47 : fragments mapped 50840
[ INFO] 2016-08-20 10:27:47 : p fragments mapped 0.89
[ INFO] 2016-08-20 10:27:47 : good mappings 8276
[ INFO] 2016-08-20 10:27:47 : p good mapping 0.14
[ INFO] 2016-08-20 10:27:47 : bad mappings 42564
[ INFO] 2016-08-20 10:27:47 : potential bridges 18
[ INFO] 2016-08-20 10:27:47 : bases uncovered 12751
[ INFO] 2016-08-20 10:27:47 : p bases uncovered 0.17
[ INFO] 2016-08-20 10:27:47 : contigs uncovbase 47
[ INFO] 2016-08-20 10:27:47 : p contigs uncovbase 0.78
[ INFO] 2016-08-20 10:27:47 : contigs uncovered 14
[ INFO] 2016-08-20 10:27:47 : p contigs uncovered 0.23
[ INFO] 2016-08-20 10:27:47 : contigs lowcovered 25
[ INFO] 2016-08-20 10:27:47 : p contigs lowcovered 0.42
[ INFO] 2016-08-20 10:27:47 : contigs segmented 14
[ INFO] 2016-08-20 10:27:47 : p contigs segmented 0.23
Here, the percent of good mappings is probably the first number to look at - this is mappings where both members of the pair are aligned in the correct orientation on the same contig, without overlapping either end. (See transrate metrics for more documentation.)
Using transrate to compare two transcriptomes¶
transrate can also compare an assembly to a “reference”. One nice thing about this is that you can compare two assemblies...
First, install the necessary software:
transrate --install-deps ref
Second, download a different assembly – this is one we did with the same starting reads, but without using digital normalization:
curl -O -L https://github.com/ngs-docs/2016-aug-nonmodel-rnaseq/raw/master/files/rna-assembly-nodn.fa.gz
gunzip rna-assembly-nodn.fa.gz
Compare in both directions:
transrate --assembly=rna-assembly.fa --reference=rna-assembly-nodn.fa --output=assembly-compare1
and
transrate --reference=rna-assembly.fa --assembly=rna-assembly-nodn.fa --output=assembly-compare2
First results:
[ INFO] 2016-08-20 10:35:35 : Comparative metrics:
[ INFO] 2016-08-20 10:35:35 : -----------------------------------
[ INFO] 2016-08-20 10:35:35 : CRBB hits 54
[ INFO] 2016-08-20 10:35:35 : n contigs with CRBB 54
[ INFO] 2016-08-20 10:35:35 : p contigs with CRBB 0.9
[ INFO] 2016-08-20 10:35:35 : rbh per reference 0.9
[ INFO] 2016-08-20 10:35:35 : n refs with CRBB 32
[ INFO] 2016-08-20 10:35:35 : p refs with CRBB 0.53
[ INFO] 2016-08-20 10:35:35 : cov25 18
[ INFO] 2016-08-20 10:35:35 : p cov25 0.3
[ INFO] 2016-08-20 10:35:35 : cov50 18
[ INFO] 2016-08-20 10:35:35 : p cov50 0.3
[ INFO] 2016-08-20 10:35:35 : cov75 18
[ INFO] 2016-08-20 10:35:35 : p cov75 0.3
[ INFO] 2016-08-20 10:35:35 : cov85 16
[ INFO] 2016-08-20 10:35:35 : p cov85 0.27
[ INFO] 2016-08-20 10:35:35 : cov95 14
[ INFO] 2016-08-20 10:35:35 : p cov95 0.23
[ INFO] 2016-08-20 10:35:35 : reference coverage 0.24
Second results:
[ INFO] 2016-08-20 10:36:45 : Comparative metrics:
[ INFO] 2016-08-20 10:36:45 : -----------------------------------
[ INFO] 2016-08-20 10:36:45 : CRBB hits 45
[ INFO] 2016-08-20 10:36:45 : n contigs with CRBB 45
[ INFO] 2016-08-20 10:36:45 : p contigs with CRBB 0.75
[ INFO] 2016-08-20 10:36:45 : rbh per reference 0.75
[ INFO] 2016-08-20 10:36:45 : n refs with CRBB 31
[ INFO] 2016-08-20 10:36:45 : p refs with CRBB 0.52
[ INFO] 2016-08-20 10:36:45 : cov25 17
[ INFO] 2016-08-20 10:36:45 : p cov25 0.28
[ INFO] 2016-08-20 10:36:45 : cov50 17
[ INFO] 2016-08-20 10:36:45 : p cov50 0.28
[ INFO] 2016-08-20 10:36:45 : cov75 16
[ INFO] 2016-08-20 10:36:45 : p cov75 0.27
[ INFO] 2016-08-20 10:36:45 : cov85 15
[ INFO] 2016-08-20 10:36:45 : p cov85 0.25
[ INFO] 2016-08-20 10:36:45 : cov95 15
[ INFO] 2016-08-20 10:36:45 : p cov95 0.25
[ INFO] 2016-08-20 10:36:45 : reference coverage 0.14
In this case you can see that our first assembly “covers” more of the other assembly than the other assembly does ours (rbh per reference, and reference coverage). However, you can also see that the assemblies differ quite a bit (for reasons that I haven’t tracked down).
Merging two (or more) assemblies¶
Finally, you can also use transrate to merge contigs from multiple assemblies, if you’ve used read mapping –
transrate --assembly=rna-assembly.fa \
--merge-assemblies=rna-assembly-nodn.fa \
--left=left.fq --right=right.fq \
--output=transrate-merge
...although for our assemblies here it doesn’t really improve them.
Back to index: 2016 / Aug / mRNAseq on non-model organisms
LICENSE: This documentation and all textual/graphic site content is licensed under the Creative Commons - 0 License (CC0) -- fork @ github. Presentations (PPT/PDF) and PDFs are the property of their respective owners and are under the terms indicated within the presentation.